In the field of bioinformatics, the identification of promoter regions is an important research direction. Although many tools for predicting promoters have been proposed, the reliability of these tools still needs improvement.
The authors present a deep learning model called DeePromoter, designed to analyze features of short eukaryotic promoter sequences and accurately identify promoter sequences in humans and mice.
DeePromoter combines two deep learning models, Convolutional Neural Networks (CNN) and Long Short-Term Memory Networks (LSTM).
Additionally, instead of using non-promoter regions of the genome as negative samples, the authors construct a more challenging negative sample set from promoter sequences. The reconstruction method of this negative sample set improves discriminative power and significantly reduces the number of false-positive predictions.
TATA-box是一个启动子亚序列,向其他分子指示转录开始的位置。它被命名为"TATA-box",因为其序列以重复的T和A碱基对(TATAAA)为特征。在人类基因中,有24%的基因启动子区域包含TATA-box。在真核生物中,TATA-box位于转录起始点(TSS)的上游约25个碱基对。它能够定义转录的方向,并指示要读取的DNA链。称为转录因子的蛋白质结合到包括TATA-box在内的几个非编码区域,并招募一种称为RNA聚合酶的酶,该酶从DNA合成RNA。
启动子预测的几种传统方法:
基于信号的方法:这种方法侧重于与RNA聚合酶结合位点相关的启动子元件。
包括 1.使用TATA框的特征提取和转录因子结合位点的加权矩阵进行分类;2.启动器2.0,使用不同盒子(如TATA盒、CAAT盒和GC-Box)传递给人工神经网络进行分类。
缺点是预测精度相对较弱,因为它们可能忽略序列的非元件部分。
基于内容的方法:
这种方法依赖于通过在序列上运行固定长度的窗口来计数k聚体的频率。包括 1.使用六聚体频率进行启动子预测;2.通过扫描定义为可变长度基序的特定特征,识别含有启动子的区域。缺点是这些方法可能忽略了序列中碱基对的空间信息。
基于GpG的方法:
这种方法利用GpG岛的位置,因为人类基因中的启动子区域或第一外显子区域通常含有GpG岛。缺点是只有约60%的启动子含有GpG岛,因此这类预测因子的准确性从未超过60%。
以前的工作,通过从基因组非启动子区域随机选择片段构建阴性集。这种方法是不完全合理的,因为如果正负集合之间没有交集。模型将很容易找到基本特征来区分这两个类。
基本思想:当正类(启动子)和负类之间的特征是共有的时,模型可能会在决策中忽视或减少对这些共有特征的依赖,而更多地关注其他不太明显的特征。
重构负类集合:对于每个正序列,生成一个负序列。将正序列分成20个子序列。随机选择12个子序列并进行随机替换。其余的8个子序列保持不变。
生成新的非启动子序列:应用上述重构过程到正集上,生成新的非启动子序列。
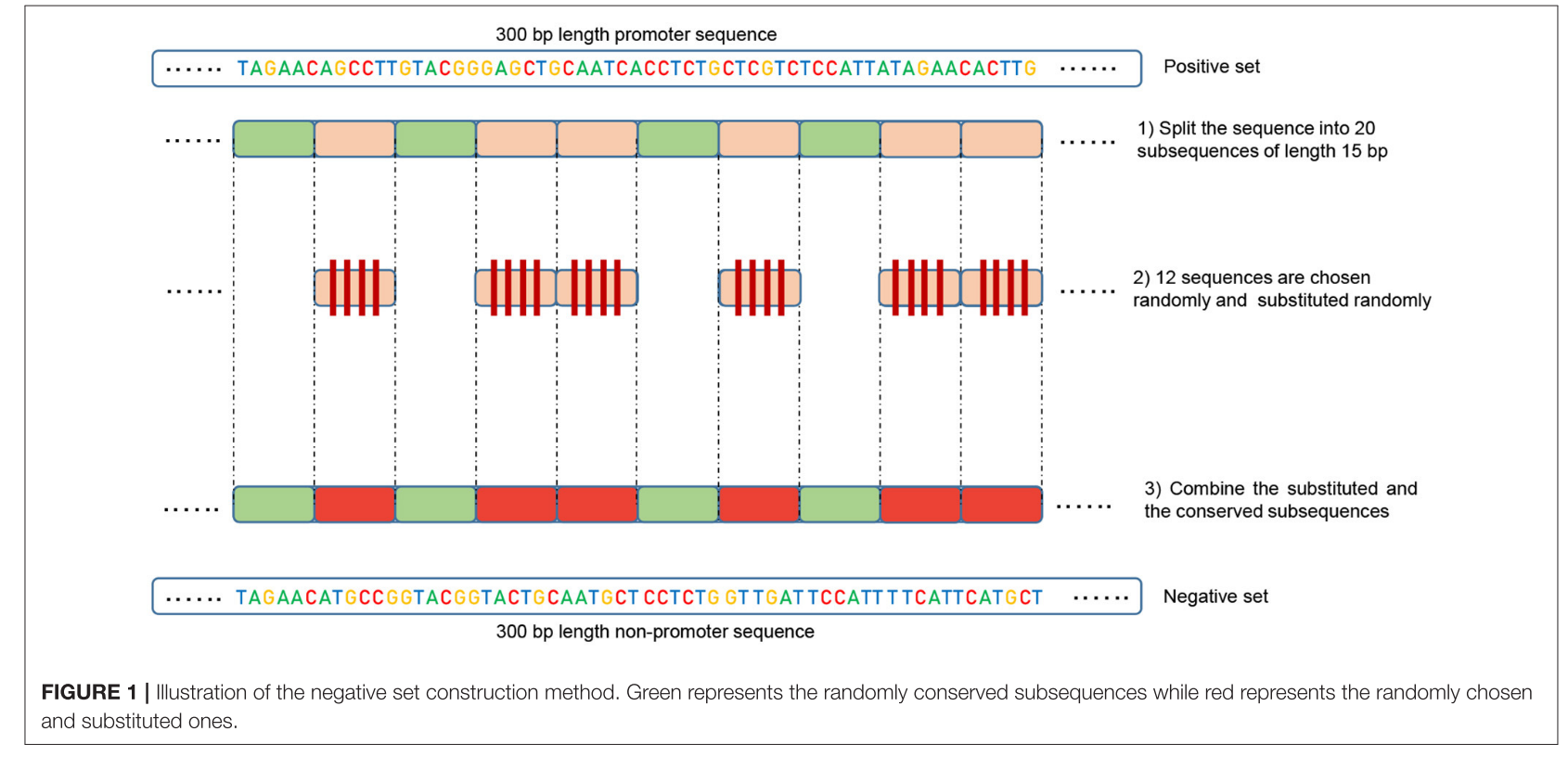
由于保守的部分在负序列中占据相同的位置,正类和负类现在共享相同的明显特征(例如,TATA-box和TSS的位置),比例为32%到40%。

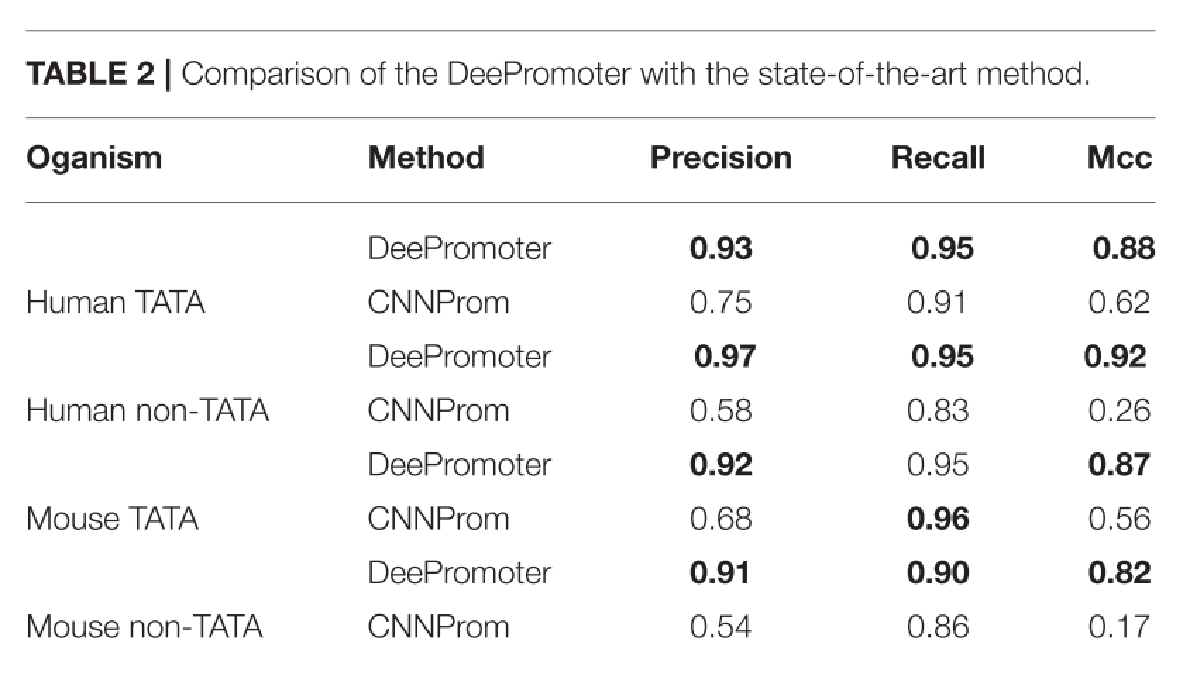
之前的工作的性能在很大程度上取决于负数据集的准备方式。它们在它们准备的数据集上表现得非常好,然而,在评估包括与启动子序列具有共同模体的更具挑战性的数据集时,它们的假阳性率较高,其中包括非启动子序列。他们的模型在处理含有TATA基序的负序列(硬例子)时严重失败。随着假阳性率的增加,精确度下降。简单地说,他们将这些序列归类为阳性启动子序列。
所提出的模型从并行对齐的多个卷积层开始,并帮助学习具有不同窗口大小的输入序列的重要图案。
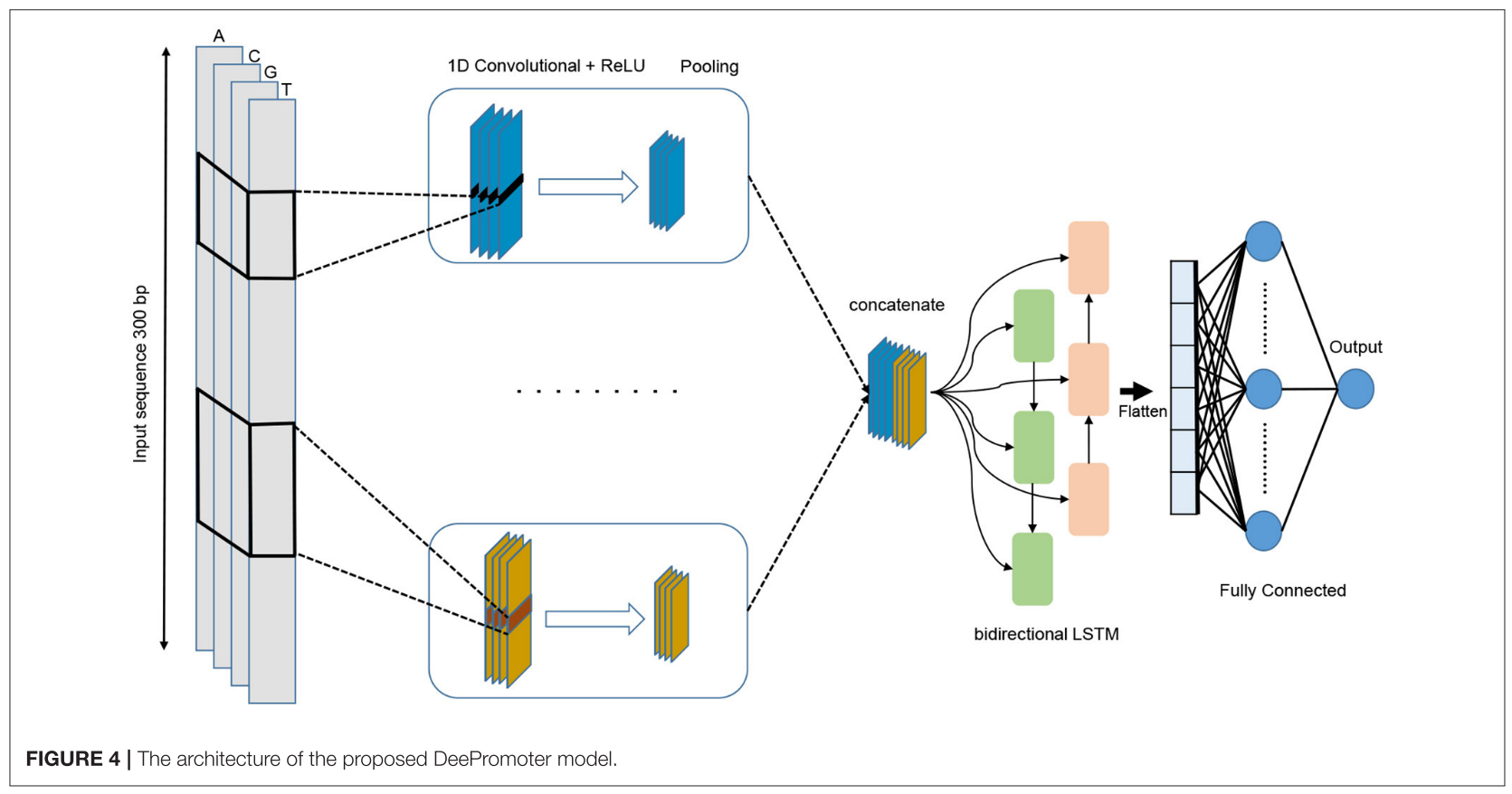
在这项工作中,我们开发了DeePromoter -它基于卷积神经网络和双向LSTM的组合-来预测人类和小鼠TATA和非TATA启动子的短真核生物启动子序列。这项工作的重要组成部分是克服在先前开发的工具中注意到的低精度(高假阳性率)的问题,这是由于在对启动子和非启动子序列进行分类时依赖于序列中的一些明显特征/基序。在这项工作中,我们特别感兴趣的是构建一个硬负集,驱动模型探索序列的深层和相关特征,而不是仅仅根据一些功能基序的存在来区分启动子和非启动子序列。
代码实现。数据加载:
import torch
import random
import numpy as np
from torch.utils.data import Dataset, DataLoader, random_split, ConcatDataset
from utils import neggen, reader, get_list_kmer, protein2num
class LoadOnehot(Dataset):
def __init__(self, pathpos, is_pos=True, device="cuda", fake=0,
length_pro=300, divide=20, part=8):
"""
Dataset
:param pathpos: txt 数据文件的路径
:param is_pos: 控制数据集的标签:True 表示 1,False 表示 0
:param device: 设备
:param fake: 0 表示加载原始 TXT 数据集,1 表示随机伪造,2 表示论文中描述的伪造方法
:param length_pro: 输入序列长度
:param divide: 在用随机序列替换某些部分之前将启动子分解成的部件数
:param part: 做随机子序列时要保持相同的部件数量
"""
if is_pos and fake != 0:
raise Exception("不能在正数据集上使用关键字fake")
self.device = device
self.fake = fake
self.length_pro = length_pro
self.divide = divide
self.part = part
# get list of kmer
dic = get_list_kmer(1)
# read data from file
self.dpos = reader(pathpos)
# convert protein to number sequence
self.npos = [protein2num(pro, dic) for pro in self.dpos]
if is_pos:
self.poslabel = torch.from_numpy(np.ones(len(self.dpos)))
else:
# ic("go false")
self.poslabel = torch.from_numpy(np.zeros(len(self.dpos)))
self.poslabel = self.poslabel.to(device)
def __len__(self):
return len(self.dpos)
def __getitem__(self, idx):
# convert data to one hot format and up to device
#截断或者填充序列长度
pro = self.npos[idx]
if len(pro) < self.length_pro:
pro = pro + [0] * (self.length_pro - len(pro))
elif len(pro) > self.length_pro:
pro = pro[:self.length_pro]
# 从原始数据随机打乱生成负样本数据集
if self.fake == 1:
pro = random.shuffle(pro)
# 通过从正样本随机替换一部分生成负样本数据集
elif self.fake == 2:
pro = neggen(pro, num_part=self.divide, keep=self.part)
torchpro = torch.from_numpy(np.array(pro))
#onehot = torch.nn.functional.one_hot(torchpro, num_classes=4).to(self.device)
onehot = torch.nn.functional.one_hot(torchpro.to(torch.int64), num_classes=4).to(self.device)
return onehot.float(), self.poslabel[idx]
def load_data(data_path, train_potion=0.8, rand_neg=False, batch_size=32, num_cpu=0, device="cuda"):
"""
加载所有数据
:param data_path: txt 文件包含启动子的路径(1 行上有 1 个 DNA 启动子)
:param train_potion: 数据集用于训练的比例
:param rand_neg: 将随机的 DNA 添加到阴性 datset 中
:param batch_size: 批量大小
:param num_cpu: 在 prallel 中执行加载数据的 CPU 数量
:param device: 将数据加载到哪个设备上
:return: 正负数据集的训练、值、测试数据集列表
"""
# 得到 dataset
manual_seed = torch.Generator().manual_seed(42)
pos_data = LoadOnehot(data_path, device=device)
neg_data = LoadOnehot('data/human/neg.txt', is_pos=False,device=device)
# 计算训练集和测试集的大小
train_num = int(len(pos_data)*train_potion)
val_num = int(len(pos_data)*(1-train_potion)*0.5)
split_size = [train_num, val_num, len(pos_data) - train_num - val_num]
# 划分数据集
train_pos, val_pos, test_pos = random_split(pos_data, split_size, generator=manual_seed)
train_neg, val_neg, test_neg = random_split(neg_data, split_size, generator=manual_seed)
# add random dataset to negative dataset(only to train set)
if rand_neg:
neg_data_rand = LoadOnehot(data_path, is_pos=False, fake=1, device=device)
train_neg = ConcatDataset([train_neg, neg_data_rand])
# data loader
stack_dataset = [train_pos, val_pos, test_pos, train_neg, val_neg, test_neg]
stack_loaders = list()
for dataset in stack_dataset:
data_loader = DataLoader(dataset, batch_size=batch_size, shuffle=True, num_workers=num_cpu)
stack_loaders.append(data_loader)
return stack_loaders
def load_data_test(data_path, batch_size=32, device="cuda", num_cpu=0):
dataset = LoadOnehot(data_path, device=device)
data_loader = DataLoader(dataset, batch_size=batch_size, shuffle=False, num_workers=num_cpu)
return data_loader
模型架构:
import random
import torch
import numpy as np
from torch import nn
from torch.nn.utils.rnn import pad_sequence
from torch.utils.data import Dataset, DataLoader, random_split
from torchvision import transforms, utils
class ParallelCNN(nn.Module):
def __init__(self, para_ker, pool_kernel=6, drop=0.5):
"""
多个 CNN 层应用于输入并连接输出,这个模型实现了一个并行的 CNN 结构,通过在输入上应用多个不同内核大小的 CNN 层,
可以提取不同尺寸的特征。这种并行结构可以增强模型对输入数据的表示能力
:param para_ker: 将使用的内核大小列表
:param pool_kernel: CNN 之后的池化参数
:param drop: Dropout 参数
"""
super(ParallelCNN, self).__init__()
self.lseq = nn.ModuleList()
for k in para_ker:
seq = nn.Sequential(
nn.Conv1d(4, 4, kernel_size=k, padding="same"),
nn.ReLU(),
nn.MaxPool1d(pool_kernel),
nn.Dropout(drop)
)
self.lseq.append(seq)
def forward(self, inputs):
"""
:param inputs: DNA onehot 序列 [batch_size x 4 x 长度]
:return: 不同内核大小的堆栈 CNN 输出功能 [batch_size x 12 x 长度]
"""
_x = list()
for seq in self.lseq:
x = seq(inputs)
_x.append(x)
# 将每个 CONV 层的输出连接到张量
_x = torch.cat(_x, 1)
return _x
class BidirectionalLSTM(nn.Module):
"""这个模型的目的是通过双向 LSTM 层对输入序列进行处理,从而获取上下文信息,并通过线性层映射到最终的输出特征"""
def __init__(self, input_size, hidden_size, output_size):
super(BidirectionalLSTM, self).__init__()
self.rnn = nn.LSTM(input_size, hidden_size, bidirectional=True, batch_first=True)
self.linear = nn.Linear(hidden_size * 2, output_size)
def forward(self, inputs):
"""
:param inputs: visual feature [batch_size x T x input_size]
:return: contextual feature [batch_size x T x output_size]
"""
self.rnn.flatten_parameters()
recurrent, _ = self.rnn(inputs) # batch_size x T x input_size -> batch_size x T x (2*hidden_size)
output = self.linear(recurrent) # batch_size x T x output_size
return output
class DeePromoter(nn.Module):
def __init__(self, para_ker, input_shape=(64, 300, 4), pool_kernel=6, drop=0.5):
"""
Deepromoter
:param para_ker: 将使用的内核大小列表
:param input_shape: 指定模型的输入形状(固定)
:param pool_kernel: CNN 之后的池化参数
:param drop: Dropout parameter
"""
super(DeePromoter, self).__init__()
binode = len(para_ker) * 4
self.pconv = ParallelCNN(para_ker, pool_kernel, drop)
self.bilstm = BidirectionalLSTM(binode, binode, binode)
self.flatten = nn.Flatten()
x = torch.zeros(input_shape)
shape = self.get_feature_shape(x)
self.fc = nn.Sequential(
nn.Linear(shape, shape),
nn.ReLU(),
nn.Linear(shape, 2),
nn.Softmax(dim=1)
)
def get_feature_shape(self, x):
"""传递虚拟输入以查找形状
后平整层用于线性层构建"""
x = x.permute(0, 2, 1)
x = self.pconv(x)
x = x.permute(0, 2, 1)
x = self.bilstm(x)
x = self.flatten(x)
return x.shape[1]
def forward(self, x):
x = x.permute(0, 2, 1)
x = self.pconv(x)
x = x.permute(0, 2, 1)
x = self.bilstm(x)
x = self.flatten(x)
x = self.fc(x)
return x
训练文件:
import torch
import math
import argparse
import torch.optim as optim
from torch import nn
from icecream import ic
from pathlib import Path
from dataloader import load_data
from modules.deepromoter import DeePromoter
from test import evaluate, mcc
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
def train(data_path, pretrain=None, exp_name="test", training=True, ker=None, epoch_num=1000):
"""
Training
:param data_path: txt 数据文件的路径
:param pretrain: 继续训练的权重路径
:param exp_name: 用于保存结果的文件夹名称
:param training: 如果为 False,则仅执行测试
:param ker: 应用于启动子序列的列表 CNN 的内核大小
:param epoch_num: 要训练的最大轮数
"""
if ker is None:
ker = [27, 14, 7]
# create the experiment folder to save the result
output = Path("./output")
output.mkdir(exist_ok=True)
exp_folder = output.joinpath(exp_name)
exp_folder.mkdir(exist_ok=True)
# load data
ic("Data loading")
data = load_data(data_path, device=device)
train_pos, val_pos, test_pos, train_neg, val_neg, test_neg = data
# model define
net = DeePromoter(ker)
net.to(device)
# load pre-train model
if pretrain is not None:
net.load_state_dict(torch.load(pretrain))
# define loss, optimizer
criterion = nn.CrossEntropyLoss()
# optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
optimizer = optim.Adam(net.parameters(), lr=0.00001)
running_loss = 0
best_mcc = 0
best_precision = 0
best_recall = 0
break_after = 10
last_update_best = 0
pbar = range(epoch_num)
ic("Start training")
if training:
for epoch in pbar:
for i, (batch_pos, batch_neg) in enumerate(zip(train_pos, train_neg)):
inputs = torch.cat((batch_pos[0], batch_neg[0]), dim=0)
labels = torch.cat((batch_pos[1], batch_neg[1]), dim=0)
# zero the parameter gradients
optimizer.zero_grad()
# pass model to
outputs = net(inputs)
loss = criterion(outputs, labels.long())
loss.backward()
optimizer.step()
running_loss += loss.item()
if epoch % 10 == 0:
torch.save(net.state_dict(), str(exp_folder.joinpath("epoch_" + str(epoch) + ".pth")))
net.eval()
eval_data, _ = evaluate(net, [val_pos, val_neg])
precision, recall, MCC = mcc(eval_data)
net.train()
ic("Epoch :", epoch, "Experiment :", exp_name)
ic("precision :", precision)
ic("recall :", recall)
ic("MCC :", MCC)
# save best model
if precision > best_precision:
best_precision = precision
ic("Update best precision")
torch.save(net.state_dict(), str(exp_folder.joinpath("best_precision.pth")))
if recall > best_recall:
best_recall = recall
ic("Update best recall")
torch.save(net.state_dict(), str(exp_folder.joinpath("best_recall.pth")))
if MCC > best_mcc:
ic("Update best MCC")
best_mcc = MCC
torch.save(net.state_dict(), str(exp_folder.joinpath("best_mcc.pth")))
last_update_best = 0
else:
last_update_best += 1
if last_update_best >= break_after:
break
if last_update_best >= break_after:
break
# test
best_model = str(exp_folder.joinpath("best_mcc.pth"))
net.load_state_dict(torch.load(best_model))
net.eval()
eval_data, _ = evaluate(net, [test_pos, test_neg])
precision, recall, MCC = mcc(eval_data)
ic("precision :", precision)
ic("recall :", recall)
ic("MCC :", MCC)
with open(str(exp_folder.joinpath("log.txt")), "w") as f:
f.write(f"Test precision: {precision}\n")
f.write(f"Test recall: {recall}\n")
f.write(f"Test MCC : {MCC}\n")
if __name__ == '__main__':
parser = argparse.ArgumentParser()
parser.add_argument("-d", "--data", type=str,
required=True, help="path to dataset(txt file)")
parser.add_argument("-e", "--experiment_name", type=str,
default="test", help="name of folder to save output in ./output")
parser.add_argument("-w", "--weight", type=str, help="Path to pre-train")
parser.add_argument("--test", action="store_true", help="Add this flag to do test only")
args = parser.parse_args()
training = True
if args.test:
training = False
train(args.data, pretrain=args.weight, exp_name=args.experiment_name, training=training)


.png)